If you have any questions or doubts, please don’t hesitate to contact riccardo.piccinno@unipv.it.
If interested, fill and send the form for the applycation. There you will include the project(s) you are interested in, your degree program (Bachelor’s or Master’s), the motivations behind your choice, and any relevant skills or experience you may have.
Important note: most projects are also open to students who do not yet have specific prior experience. Feel free to highlight the absence of certain skills if this is motivated by a genuine desire to develop them during the internship.
After your request is received, a short introductory meeting will be scheduled to get to know each other and discuss the opportunity.
Below you will find a list of currently available projects. Start dates are indicative. When the “type of internship” includes more than one category, this means that the project can be approached from multiple perspectives, it does not imply that the student is expected to cover all aspects. The specific focus will be defined together, based on your interests and the discussion during the initial meeting.
Students are also warmly encouraged to propose their own ideas. Their feasibility will be assessed and discussed together during the interview. Apply here with your ideas.
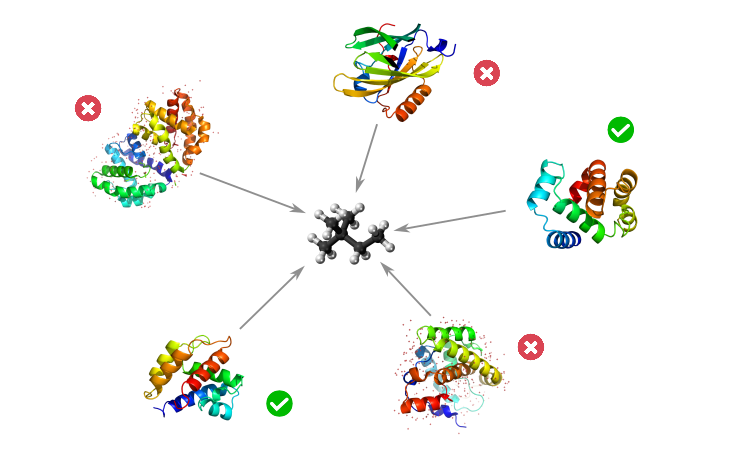
Design and development of ProteinSearch, a pipeline to automate the search for proteins that interact with a given molecule in silico.
For more information, contact riccardo.piccinno@unipv.it.
- Project start: From 01/12/2025
- Suitable for: Master’s students
- Internship type: Computational
- Required skills: Basic knowledge of Bash and Python. Familiarity with PyMOL is appreciated.
- Skills acquired: Design and implementation of bioinformatics pipeline
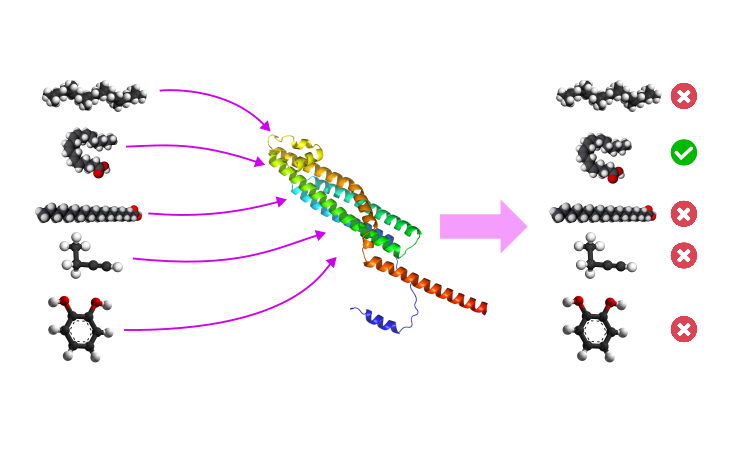
Design and development of LigMS, a pipeline to automate the search for ligands for a given protein in silico.
For more information, contact riccardo.piccinno@unipv.it.
- Project start: From 01/12/2025
- Suitable for: Master’s students
- Internship type: Computational
- Required skills: Basic knowledge of Bash and Python. Familiarity with PyMOL is appreciated.
- Skills acquired: Design and implementation of bioinformatics pipeline

Design and development of consequence, a bioinformatics tool aimed at simplifying and making high-quality phylogenetic analysis more accessible. For further details, please visit consequence or contact riccardo.piccinno@unipv.it.
- Project start: ASAP
- Suitable for: Bachelor’s/Master’s students
- Internship type: Computational
- Required skills: None. Familiarity with Python and Bash is appreciated
- Skills acquired: Design and implementation of bioinformatics tools and pipelines
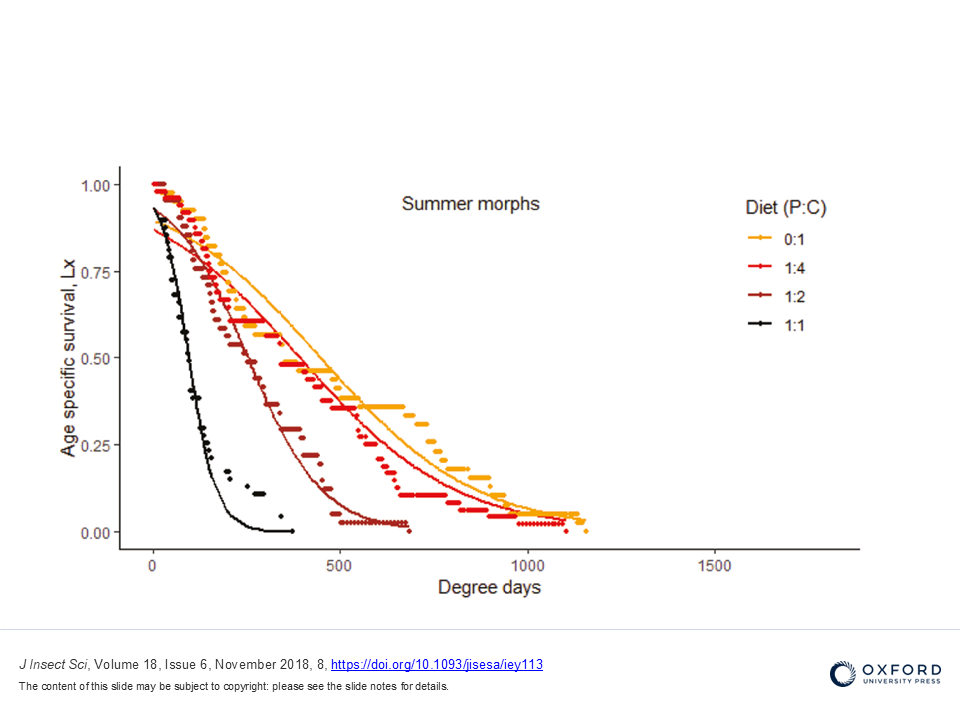
Development of survival models to estimate the impact of Wolbachia infection on the ability of D. suzukii to overwinter. For more information, visit WICKED or contact riccardo.piccinno@unipv.it.
- Project start: From 01/04/2026
- Suitable for: Master’s students
- Internship type: Laboratory and/or Computational
- Required skills: None. Experience in insect handling, PCR, R programming and/or statistics is a plus but not required
- Skills acquired: Understanding and development of biostatistical models and data analysis
- Related publications:
- Perlmutter, J. I., Atadurdyyeva, A., Schedl, M. E., & Unckless, R. L. (2025). Wolbachia enhances the survival of Drosophila infected with fungal pathogens. BMC biology, 23(1), 42. https://doi.org/10.1186/s12915-025-02130-0
- Adonyeva, N. V., Efimov, V. M., & Gruntenko, N. E. (2023). The Effect of Genotype Combinations of Wolbachia and Its Drosophila melanogaster Host on Fertility, Developmental Rate and Heat Stress Resistance of Flies. Insects, 14(12), 928. https://doi.org/10.3390/insects14120928
- Panel, A. D. C., Pen, I., Pannebakker, B. A., Helsen, H. H. M., & Wertheim, B. (2020). Seasonal morphotypes of Drosophila suzukii differ in key life-history traits during and after a prolonged period of cold exposure. Ecology and evolution, 10(17), 9085–9099. https://doi.org/10.1002/ece3.6517
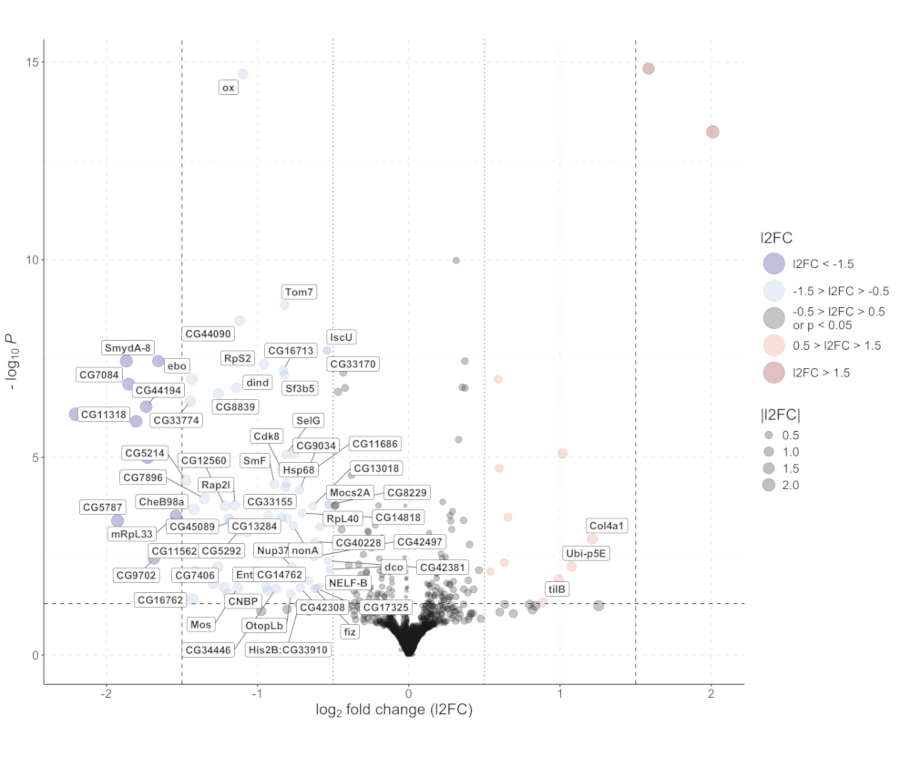
Design and analysis of RNAseq experiments to study the effects of Wolbachia infection on gene expression and microbiota composition in D. suzukii. For further details, refer to WICKED or contact riccardo.piccinno@unipv.it.
- Project start: From 01/04/2026
- Suitable for: Master’s students
- Internship type: Laboratory and/or Computational
- Required skills: For the wet-lab component, lab tool proficiency is required; prior experience in nucleic acid extraction is appreciated. For the computational component, knowledge of R is required; familiarity with Bash and nf-core pipelines is a plus
- Skills acquired: Wet-lab: RNA extraction, rRNA depletion, RT-PCR/dPCR. Computational: use of HPC resources, Bash scripting, differential gene expression and metatranscriptomic analysis
- Related publications:
- Enriquez, T., & Colinet, H. (2019). Cold acclimation triggers major transcriptional changes in Drosophila suzukii. BMC genomics, 20(1), 413. https://doi.org/10.1186/s12864-019-5745-7
- Plantamp, C., Henri, H., Andrieux, T., Régis, C., Mialdea, G., Dray, S., Gibert, P., & Desouhant, E. (2019). Phenotypic plasticity in the invasive pest Drosophila suzukii: activity rhythms and gene expression in response to temperature. The Journal of experimental biology, 222(Pt 14), jeb199398. https://doi.org/10.1242/jeb.199398
- Shearer, P. W., West, J. D., Walton, V. M., Brown, P. H., Svetec, N., & Chiu, J. C. (2016). Seasonal cues induce phenotypic plasticity of Drosophila suzukii to enhance winter survival. BMC ecology, 16, 11. https://doi.org/10.1186/s12898-016-0070-3
- Martinez-Sañudo, I., Simonato, M., Squartini, A., Mori, N., Marri, L., & Mazzon, L. (2018). Metagenomic analysis reveals changes of the Drosophila suzukii microbiota in the newly colonized regions. Insect science, 25(5), 833–846. https://doi.org/10.1111/1744-7917.12458
Investigation of the structural basis for the malfunction of Wolbachia factors in the wSuz strain using pull-down assays and crystallography. The aim is to identify possible conformational anomalies responsible for the lack of reproductive manipulation. For more information, see WICKED or contact riccardo.piccinno@unipv.it.
- Project start: From 01/12/2025, computational done
- Suitable for: Master’s students
- Internship type: Laboratory and/or Computational
- Required skills: For the wet-lab component, lab proficiency is required; prior experience in nucleic acid extraction is appreciated. For the computational component, knowledge of Bash is required
- Skills acquired: Wet-lab: recombinant protein production, pull-down assays, crystallography. Computational: use of HPC, Bash scripting, molecular dynamics simulations
- Related publications:
- Wang, H., et al. (2022). Crystal Structures of Wolbachia CidA and CidB Reveal Determinants of Bacteria-induced Cytoplasmic Incompatibility and Rescue. Nature Communications, 13(1), 1608. https://doi.org/10.1038/s41467-022-29273-w
- Cattel, J., et al. (2018). Back and forth Wolbachia transfers reveal efficient strains to control spotted wing drosophila populations. Journal of Applied Ecology, 55(5), 2408-2418. https://doi.org/10.1111/1365-2664.13101
In collaboration with: The Armenise-Harvard Laboratory of Structural Biology – Prof. Federico Forneris
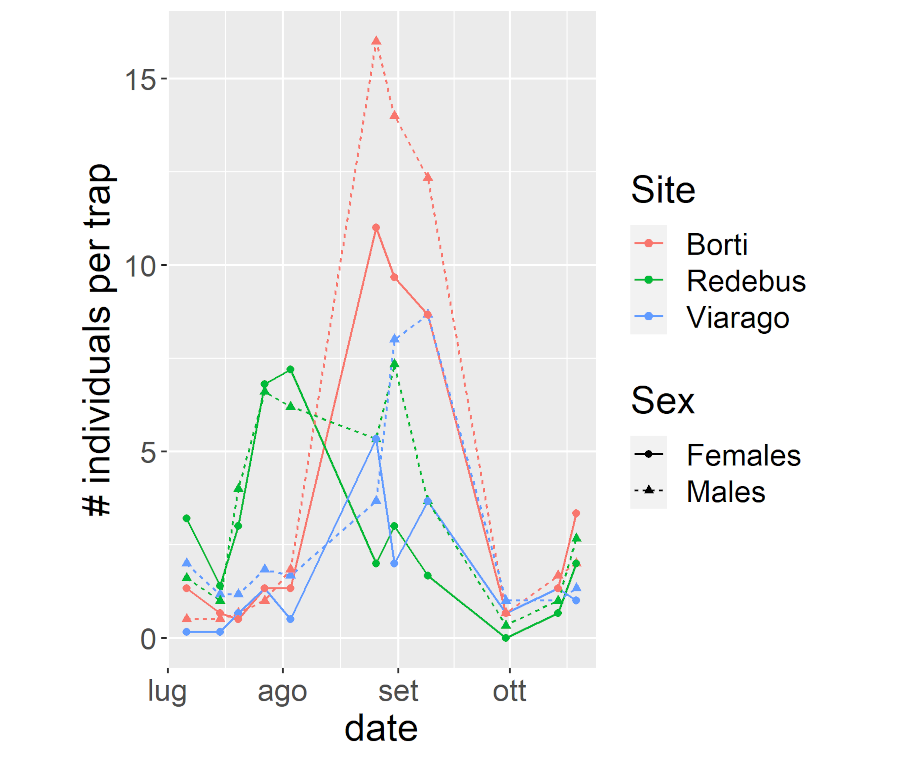
Statistical modeling aimed at estimating how the prevalence of Wolbachia varies in wild populations of D. suzukii as a function of environmental and nutritional factors. For more information, visit WICKED or contact riccardo.piccinno@unipv.it.
- Project start: From 02/02/2026
- Suitable for: Bachelor’s students
- Internship type: Laboratory and /or Computational
- Required skills: None. Experience with PCR, R programming and/or statistics is appreciated but not necessary
- Skills acquired: Development of biostatistical models and data analysis techniques
- Related publications:
- Wiman, N. G., Dalton, D. T., Anfora, G., Biondi, A., Chiu, J. C., Daane, K. M., Gerdeman, B., Gottardello, A., Hamby, K. A., Isaacs, R., Grassi, A., Ioriatti, C., Lee, J. C., Miller, B., Stacconi, M. V., Shearer, P. W., Tanigoshi, L., Wang, X., & Walton, V. M. (2016). Drosophila suzukii population response to environment and management strategies. Journal of pest science, 89, 653–665. https://doi.org/10.1007/s10340-016-0757-4
- McPherson, A. E., Abram, P. K., Curtis, C. I., Wannop, E. R., Dudzic, J. P., & Perlman, S. J. (2023). Dynamic changes in Wolbachia infection over a single generation of Drosophila suzukii, across a wide range of resource availability. Ecology and evolution, 13(11), e10722. https://doi.org/10.1002/ece3.10722
- Auguste, A., Ris, N., Belgaidi, Z., Kremmer, L., Mouton, L., & Fauvergue, X. (2024). Insect population dynamics under Wolbachia-induced cytoplasmic incompatibility: Puzzle more than buzz in Drosophila suzukii. PloS one, 19(3), e0300248. https://doi.org/10.1371/journal.pone.0300248
