About AniMEE lab
|
Numerous natural forces act on living organisms, shaping their evolution. Thanks to these selective pressures, both abiotic (such as temperature and humidity) and biotic (such as interactions with other organisms), we observe today an extraordinary biodiversity, even in the most extreme environments. The Laboratory of Animal Molecular Ecology and Evolution (AniMEE Lab) investigates how these forces drive animal evolution through an interdisciplinary approach that combines biology, molecular biology, bioinformatics, and biostatistics. Main methodological approaches include:
|
 |
At AniMEE, these methodologies are applied to questions of ecological, agricultural, veterinary, and public health relevance. The lab also develops bioinformatics tools designed to make these advanced analyses accessible to users without specialized technical expertise.
If you are a student interested in an internship, look here or apply here with your ideas.
Ecological Applications
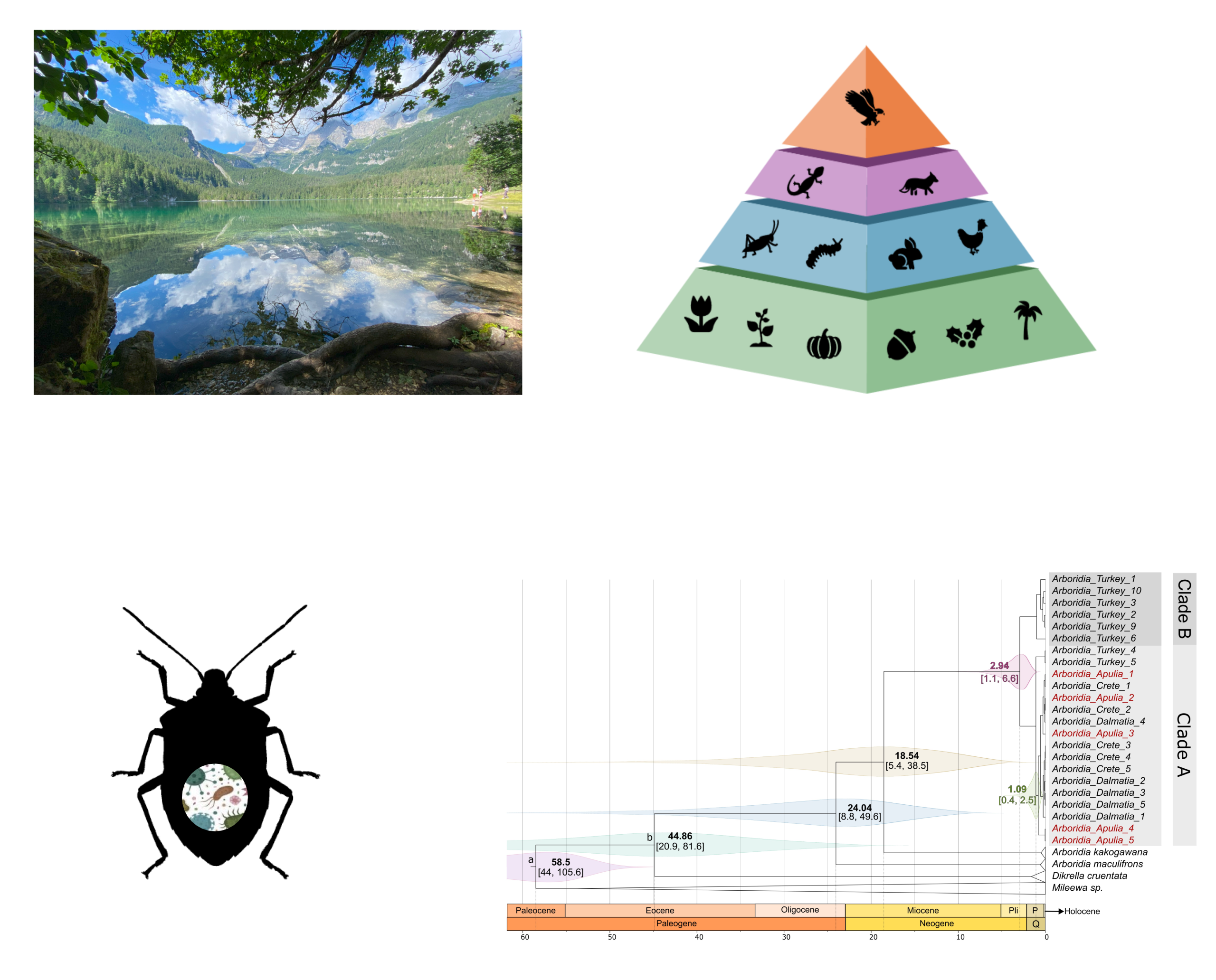 Top right: example of a food chain. Bottom left: interaction between host and microbiota. Bottom right: example of molecular dating of species origin. |
The molecular methods adopted in AniMEE are essential for biodiversity assessment and monitoring. Techniques such as barcoding, metabarcoding, and metagenomics are used to analyze environmental DNA (eDNA), providing detailed snapshots of biodiversity within ecosystems (including animals, plants, fungi, and bacteria). These tools are also applied to the study of trophic networks, enabling the reconstruction of predation dynamics and the development of predictive mathematical models. Such analyses allow for the early detection of changes invisible to direct observation, with significant implications for species conservation, invasive species monitoring and control. Phylogenetics/phylogenomics and population genetics/genomics are equally important for understanding evolutionary relationships, the impact of ecological factors on evolution, and for identifying events such as hybridization or coevolution. AniMEE is currently developing consequence, a bioinformatics tool that will enable complete evolutionary analyses through a guided, user-friendly interface. Additionally, metagenomics and metataxonomics are used to study how host–microbiota interactions vary across species or among populations of the same species. Another major area of research is pangenomics, the study of the full set of genes across populations within a taxonomic group. In this context, AniMEE is developing a project on the pan-metagenome, aimed at characterizing symbiotic microbial variability across different animal populations to explore the full spectrum of host–microbe interactions for a given species. |
Associated publications:
- Vilaça, S. T., Piccinno, R., Rota-Stabelli, O., Gabrielli, M., Benazzo, A., Matschiner, M., Soares, L. S., Bolten, A. B., Bjorndal, K. A., & Bertorelle, G. (2021). Divergence and hybridization in sea turtles: Inferences from genome data show evidence of ancient gene flow between species. Molecular ecology, 30(23), 6178–6192. https://doi.org/10.1111/mec.16113
- Piccinno, R., Tatti, A., Avosani, S., Galla, G., Lazazzara, V., Pedrazzoli, F., Zadra, N., Rodeghiero, M., Seljak, G., Özgen, İ., Hauffe, H. C., Verrastro, V., Rossi Stacconi, M. V., Mazzoni, V., & Rota-Stabelli, O. (2024). A multidisciplinary approach to tackling invasive species: barcoding, morphology, and metataxonomy of the leafhopper Arboridia adanae. Scientific reports, 14(1), 2229. https://doi.org/10.1038/s41598-023-49410-9
Agricultural Applications
|
The agricultural sector is increasingly threatened by pest species, particularly invasive ones, whose impacts are exacerbated by climate change and globalization. Notable examples include Drosophila suzukii (spotted-wing drosophila) and Halyomorpha halys (brown marmorated stink bug), both of which cause serious damage to crops due to their rapid adaptability, resistance to conventional control methods, and absence of natural enemies in newly invaded areas. In this context, studying pest ecology is crucial for identifying biological control agents, such as parasitoids or symbiotic bacteria, that can be safely introduced to manage or eradicate infestations. AniMEE’s flagship project in this field is WICKED, funded by Fondazione Cariplo, which investigates the role of the bacterium Wolbachia in D. suzukii and its potential for biological control. The project integrates biostatistics, transcriptomics, metatranscriptomics, and structural biology (in collaboration with the Armenise-Harvard Structural Biology Laboratory, led by Prof. Federico Forneris). Wolbachia is known for its ability to manipulate insect reproduction and is currently used in combination with the Sterile Insect Technique (SIT) to reduce pest populations. Associated publications:
|
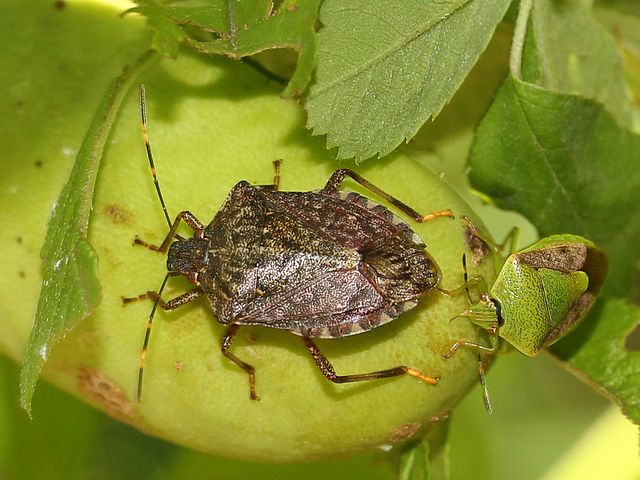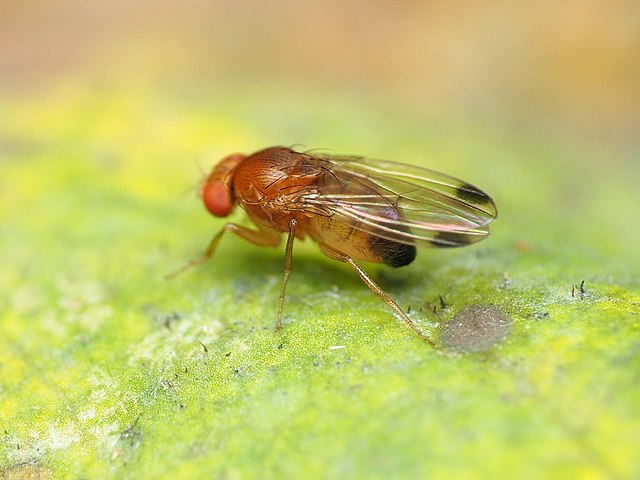  Bottom: spotted wing drosophila (Drosophila suzukii). Images sourced from Wikimedia Commons. |
Public Health and Veterinary Applications
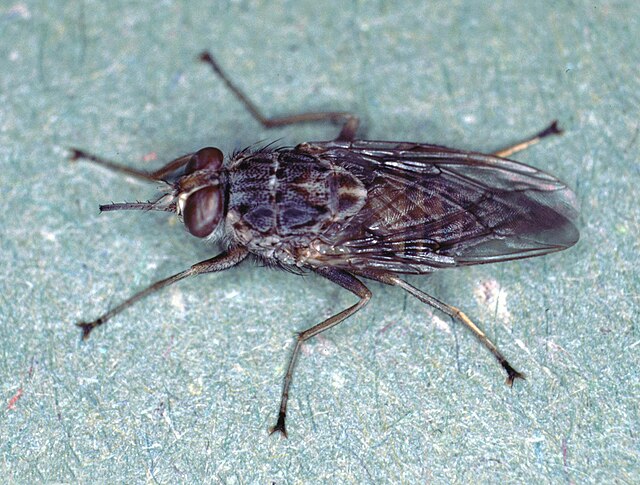  Bottom: tsetse fly (Glossina morsitans). Imaged sourced from Wikimedia Commons. |
The same molecular and bioinformatic tools used in agriculture are also applied to public health and veterinary science. Several animal species, both native and invasive, pose direct threats to humans and domestic or livestock animals as vectors of pathogens and parasites. AniMEE is currently studying the Asian tiger mosquito (Aedes albopictus) and the tsetse fly (Glossina spp.), using population genetics, phylogenetics, metataxonomics, and transcriptomics to understand the ecological and evolutionary dynamics of these vectors and to develop more effective containment strategies. Associated publications:
|